Supported Data Formats¶
About this guide
This guide provides a basic overview of file formats and data supported in the ODM. For a detailed description and instructions on using various data formats and working with them (sorting, filtering, sampling), visit the Supported Data Formats page in our Advanced User Guide.
-
Upload and manage tabular data (TSV files) seamlessly within the ODM. Work with Samples, Libraries, Preparations, Expression data, and more.
-
Upload and work with GCT (Gene Cluster Text) files in the Open Data Manager (ODM). Optimize the analysis of matrix-compatible datasets.
-
Upload and work with VCF (Variant Call Format) files to search, filter, retrieve, and analyze genetic variants in the ODM.
-
Upload and store HDF5 (Hierarchical Data Format 5) files as attachments in the ODM. Future releases will enable seamless analysis of Single Cell data stored within TSV and HDF5 formats.
-
Upload and work with FACS (Fluorescence-Activated Cell Sorting) files in the Open Data Manager (ODM) to efficiently analyze Flow Cytometry data.
-
Upload and organize a diverse range of attached non-indexed files in the ODM. Easily manage your entire data catalog, access and collaboration across any file types.
TSV (Tabular data)¶
In ODM, you can upload any tabular data formatted as TSV (tab-separated values). As long as your file represents a data frame, ODM can import and index it. A data frame is a data structure that organizes data into a two-dimensional table of rows and columns, similar to a spreadsheet.
A data frame contains two main elements:
- Features: These are the entities measured in an experiment (e.g., genes, proteins, metabolites, pathways, sales regions, etc.).
- Measurements (or values): These are the actual values recorded for each feature under different conditions (e.g., gene expression values, protein abundance, pathway activity, sales volume, etc.).
Simple Data Frame¶
The example below demonstrates the simplest and most common type of data frame.
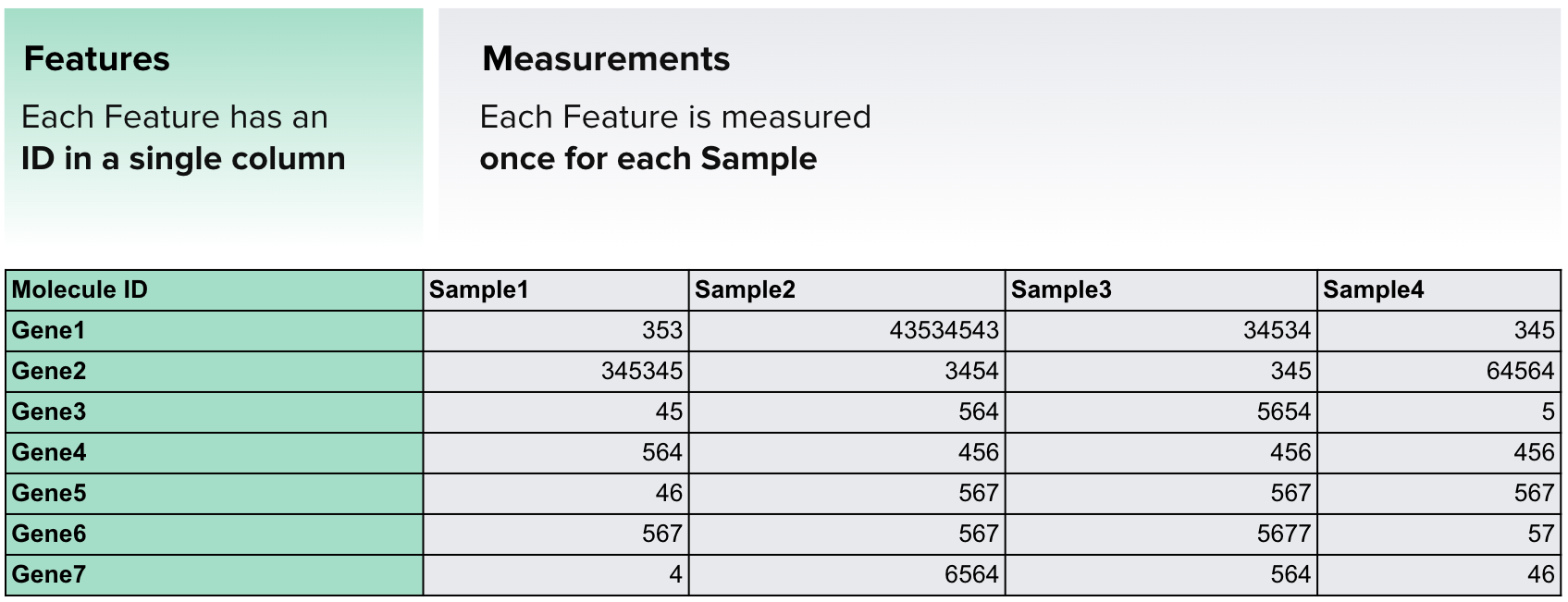
Here, the features (genes) are listed in the first column, while the rest of the table contains measurements of gene expression across multiple samples. Each column represents a different sample, with the column name indicating the corresponding dataset of gene expression values.
Complex Data Frame¶
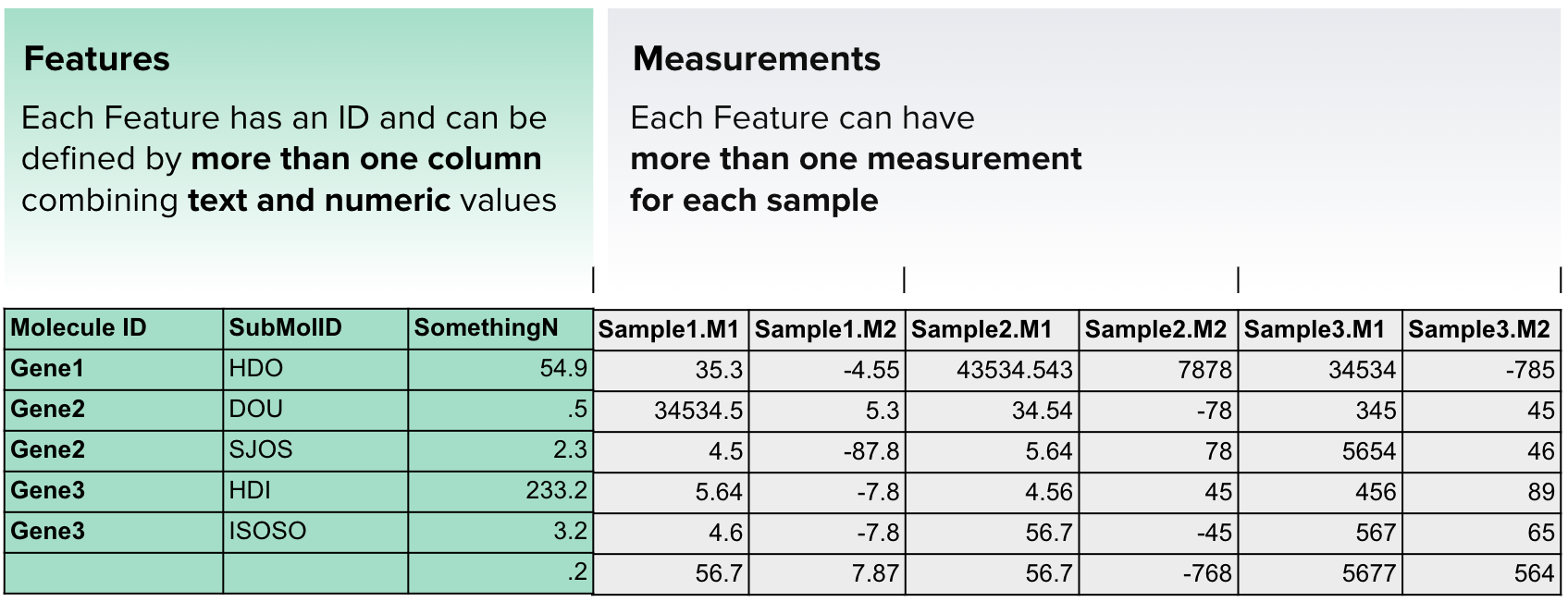
This format provides a wide range of data types that can be uploaded and indexed in ODM.
For a detailed description and instructions on using TSV, visit the Supported Data Formats page in our Advanced User Guide.
GCT (Gene Expression)¶
The ODM supports GCT (Gene Cluster Text) files, which are commonly used for storing gene expression datasets, such as microarray and RNA-seq data. These files provide a structured, tab-delimited format for organizing expression values across different samples. ODM automatically recognizes GCT files enabling integration and analysis.
Supported GCT Formats¶
ODM accepts the following GCT file formats:
- .gct – Standard GCT file
- .gct.gz, .gct.zip – Compressed versions of the standard GCT file (available via API)
- .gct.tsv, .gct.tsv.gz, .gct.tsv.zip – GCT files with additional expression metadata (available via API)
Structure of a GCT File¶
A GCT file consists of a structured matrix with gene expression values. The key components include:
- File Version: The first line always contains the file version, which is
#1.2for the GCT format. - Matrix Dimensions: The second line specifies the number of genes (rows) and the number of samples (columns), excluding metadata columns.
- Header Row: The third line contains column labels:
Name(gene identifier, case insensitive)Description(text description of the gene, case insensitive)Sample identifiers(unique, single-word names without spaces)
- Data Matrix:
- Each row corresponds to a gene, with its identifier and description in the first two columns.
- The remaining columns contain expression values for each sample.
For a detailed description and instructions on using GCT, visit the Supported Data Formats page in our Advanced User Guide.
VCF (Variants)¶
The ODM supports VCF (Variant Call Format) files, a widely used format for storing genetic variation data. VCF files provide a structured, tab-delimited representation of genetic variants and are typically generated as output from variant calling pipelines. These files contain detailed information about sequence variations, including single nucleotide polymorphisms (SNPs), insertions, deletions, and structural variants.
Supported VCF Formats¶
ODM accepts the following VCF file formats:
- .vcf – Standard uncompressed VCF file
- .vcf.gz, .vcf.zip – Compressed versions of the standard VCF file (available via API)
Structure of a VCF File¶
A VCF file consists of three main components:
- Meta-information lines (
##) - Header line (
#) – The final metadata line, which defines the column names for variant data:CHROM(chromosome)POS(genomic position)ID(variant identifier)REF(reference allele)ALT(alternate allele(s))QUAL(quality score)FILTER(filter status)INFO(additional annotations)
- Data lines – Each row represents a variant, detailing its position, reference and alternate alleles, quality scores, and annotations.
Common Fields in VCF Files¶
VCF files provide essential information for genomic studies, with key fields including:
- INFO: Contains annotations about the variant, such as depth of coverage (
DP), allele frequency (AF), and functional effect (EFF). - FILTER: Specifies whether the variant has passed quality control thresholds (
PASSor filter conditions likeq10for quality < 10). - FORMAT: Defines the structure of genotype-related fields in the sample data.
- Sample Columns: Contain individual genotype information, including genotype (
GT), depth (DP), and phasing status (PS).
For detailed specifications on the VCF format, refer to the official VCF documentation.
Using VCF Files in ODM¶
The ODM allows users to:
- Import and store genomic variants.
- Integrate variant data with sample metadata for downstream analysis.
- Searching and filtering of genetic variants.
- Perform cross-sample comparisons and study variant distributions.
For a detailed description and instructions on ODM capabilities for using VCF, visit the Supported Data Formats page in our Advanced User Guide.
HDF5 (e.g. Single Cell)¶
Limitations
HDF5 is supported as Attached File in ODM with ability to observe and search by File Structure (Contents) only. We are working on full functionality for HDF5 data content parsing, search, and filtering.
HDF5 (Hierarchical Data Format version 5) is a widely used data format in genomic research, particularly in Single Cell studies. It is designed to store large, complex datasets efficiently, making it a preferred choice for structured biological data such as gene expression matrices and metadata.
The ODM now supports HDF5 file upload as Attached File, search by File Structure (Contents), and manage these files within Studies.
Supported HDF5 Formats¶
- .h5, .h5ad - Standard HDF5 files
- .h5.gz, .h5ad.zip - Compressed versions of standard HDF5 files
Viewing File Structure (Contents)¶
- Access File Contents via GUI: The ODM displays File Contents on the Data Tab of Metadata Editor. It is
accessible on
Contentsbutton click. - Retrieve File Contents via API: You can retrieve File Contents for the list of files or by unique Genestack Accession.
Searching HDF5 Files by File Contents¶
Users can search via GUI and API for:
- Unique File: by Genestack Accession.
-
Files:
- By File Contents fields/pathways.
- By Study Genestack Accession.
-
Studies:
- By File Contents fields/pathways.
- By File Genestack Accession.
FACS (Flow Cytometry)¶
Data Overview¶
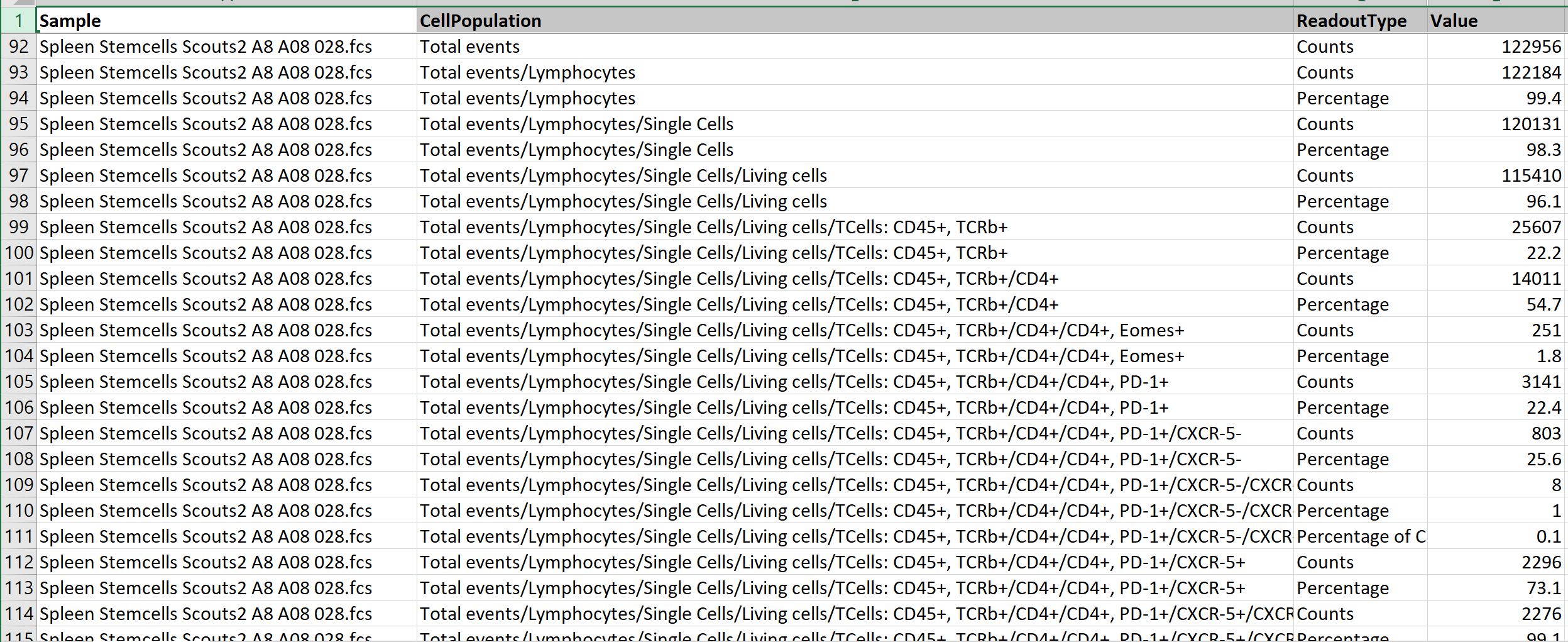
- The FACS format in the ODM represents processed (post-gating) flow cytometry data for human-readable analysis and integration.
-
Key Columns:
-
Sample:
- A string representing the sample name without additional extraction.
-
CellPopulation:
- Describes cell subtypes in a hierarchical, path-like structure (e.g.,
CD45+, live/CD45+, CD3+/CD4). - Counts at parent levels equal the sum of their children, but marker-specific counts may not add up due to cells expressing multiple markers.
- Describes cell subtypes in a hierarchical, path-like structure (e.g.,
-
ReadoutType:
- Defines the type of value recorded:
- Median: Median intensity of a fluorophore (decimal).
- Count: Number of cells (integer).
- Percentage: Proportion relative to the parent population.
-
Marker:
- Identifies proteins or fluorophores on the cell surface (e.g.,
PD-1,GZB,BV786). - Some markers may have different names but refer to the same entity.
- Identifies proteins or fluorophores on the cell surface (e.g.,
-
-
Considerations:
- Cell populations are hierarchical, and cells may carry multiple markers simultaneously.
- Marker counts do not fully represent the total population due to overlapping markers.
Using FACS in ODM¶
ODM indexes FACS files and provide API endpoints to search via them:
-
Find FACS objects and groups by its metadata and data:
- Run
- Readout Type
- Population
- Marker
- Value
-
Find objects and groups by its Genestack Accession.
-
Find objects and groups by entities:
- Study
- Samples
-
Update FACS group metadata by object id.
Attached Files¶
The ODM allows users to upload and manage any attachments related to their Studies. Attachments can be uploaded via the GUI or API, and for every uploaded file must be assigned a Data Class to ensure proper organization within catalogue and retrieval.
Uploading Attachments¶
Users can upload any file type through the GUI, the POST /api/v1/jobs/import/file endpoint in job endpoints definition, or the import_odm_data.py script.
All uploaded attachments are stored in S3 and reflected in the system but are not indexed.
To maintain structured catalogue, each attachment must be assigned a Data Class.
Limitation
- S3 bucket is mandatory to upload and work with Attached files functionality in the ODM.
- Export: If attachment's metadata was updated and got a new version, attached file cannot be exported itself from the ODM. Workaround: export is available from exporting whole Study. We are working on improvements for this functionality in 1.61 release.
Managing and Viewing Attachments¶
Attachments are displayed in the appropriate Data Class group on the Data tab in User Interface.
Users can view and manage uploaded files, ensuring they are correctly categorized and accessible within their Studies.
Metadata can be added, edited, and customized for attachments, and additional non-template attributes can be specified.
Searching for Attachments¶
The Study Browser and API allows users to find Studies by Attached file metadata. Study Browser also allows to search for Studies by Data Class assigned to Attached files.
API Support¶
The API provides endpoints to manage attachments:
- Upload Attached files.
- Find all Attached files metadata.
- Find metadata of particular Attached file.
- Find Studies by Attached file id or metadata.
- Download Attached file.
- Delete Attached files.